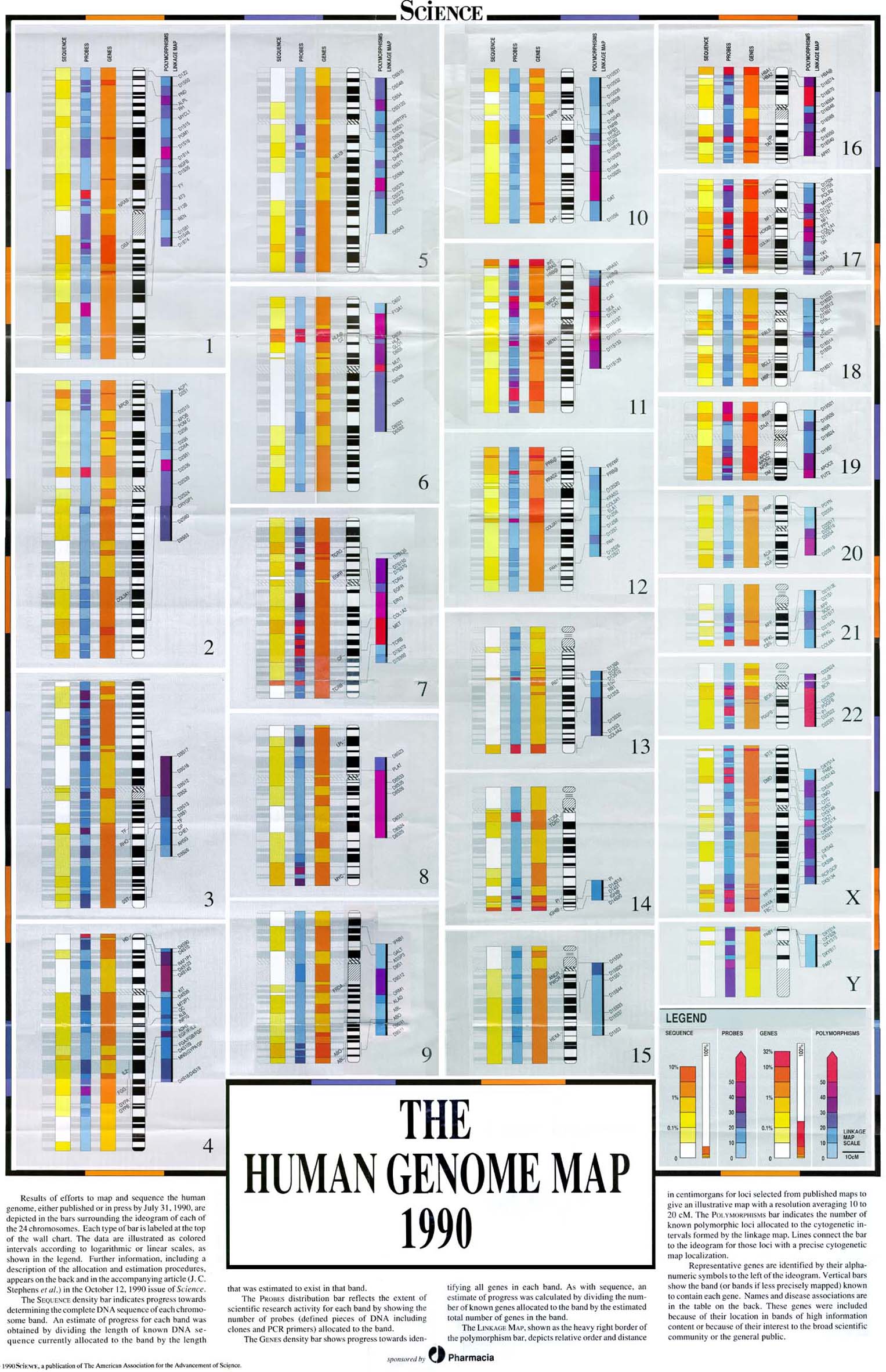
On 5 September, an international team of researchers reveal that much of what has been called ‘junk DNA’ in the human genome is actually a massive control panel with millions of switches regulating the activity of our genes. Without these switches, genes would not work – and mutations in these regions might lead to human disease. Discovered by hundreds of scientists working on the ENCODE Project, the new information is so comprehensive and complex that it has given rise to a new publishing model in which electronic documents and datasets are interconnected.

Just as the Human Genome Project revolutionised biomedical research, ENCODE will drive new understanding and open new avenues for bio-medical science. Led by the National Genome Research Institute (NHGRI) in the US and the EMBL-European Bioinformatics Institute (EMBL-EBI) in the UK, ENCODE now presents a detailed map of genome function that identifies 4 million gene ‘switches’. This essential reference will help researchers pinpoint very specific areas of research for human disease. The findings are published in 30 connected, open-access papers appearing in three science journals: Nature, Genome Biology and Genome Research.
“Our genome is simply alive with switches: millions of places that determine whether a gene is switched on or off,” says Ewan Birney of EMBL-EBI, lead analysis coordinator for ENCODE. “The Human Genome Project showed that only 2% of the genome contains genes, the instructions to make proteins. With ENCODE, we can see that around80% of the genome is actively doing something. We found that a much bigger part of the genome – a surprising amount, in fact – is involved in controlling when and where proteins are produced, than in simply manufacturing the building blocks.”
“ENCODE data can be used by any disease researcher, whatever pathology they may be interested in,” said Ian Dunham of EMBL-EBI, who played a key role in coordinating the analysis. “In many cases you may have a good idea of which genes are involved in your disease, but you might not know which switches are involved. Sometimes these switches are very surprising, because their location might seem more logically connected to a completely different disease. ENCODE gives us a set of very valuable leads to follow to discover key mechanisms at play in health and disease. Those can be exploited to create entirely new medicines, or to repurpose existing treatments.”
“ENCODE gives us the knowledge we need to look beyond the linear structure of the genome to how the whole network is connected,” commented Dr Michael Snyder, professor and chair at Stanford University and a principal investigator on ENCODE. “We are beginning to understand the information generated ingenome-wide association studies – not just where certain genes are located, but which sequences control them. Because of the complex, three-dimensional shape of our genome, those controls are sometimes far from the gene they regulate and looping around to make contact. Were it not for ENCODE, we might never have looked in those regions. This is a major step toward understanding the wiring diagram of a human being. ENCODE helps us look deeply into the regulatory circuit that tells us how all of the parts come together to make a complex being.”
Until recently, generating and storing large volumes of data has been a challenge in biomedical research. Now, with the falling cost and rising productivity of genome sequencing, the focus has shifted to analysis – making sense of the data produced in genome-wide association studies. ENCODE partners have beenworking systematically through the human genome, using the same computational and wet-lab methods and reagents in laboratories distributed throughout the world.
To give some sense of the scale of the project: ENCODE combined the efforts of 442 scientists in 32 labs in the UK, US, Spain, Singapore and Japan. They generated and analysed over 15 terabytes (15 trillion bytes) of raw data – all of which is now publicly available. The study used around 300 years’ worth of computer time studying 147 tissue types to determine what turns specific genes on and off, and how that ‘switch’ differs between cell types.
The articles published today represent hundreds of pages of research. But the digital publishing group atNature recognizes that ‘pages’ are a thing of the past. All of the published ENCODE content, in all three journals, is connected digitally through topical ‘threads’, so that readers can follow their area of interest between papers and all the way down to the original data.
“Getting the best people with the best expertise together is what this is all about,” said Ewan Birney. “ENCODE has really shown that leading life scientists are very good at collaborating closely on a large scale to produce excellent foundational resources that the whole community can use.”
“Until now, everyone’s been generating and publishing this data piecemeal and unintentionally trapping it in niche communities and static publications. How could anyone outside that community exploit that knowledge if they don’t know it’s there?” commented Roderic Guigo of the Centre de Regulació Genómica (CRG) in Barcelona, Spain. “We have now an interactive encyclopedia that everyone can refer to, and that will make a huge difference.”
#This press release is copied from http://www.ebi.ac.uk/